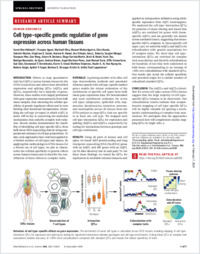
Cell type-specific genetic regulation of gene expression across human tissues.
- Kim-Hellmuth S Statistical Genetics, Max Planck Institute of Psychiatry, Munich, Germany. skimhellmuth@gmail.com tlappalainen@nygenome.org.
- Aguet F The Broad Institute of MIT and Harvard, Cambridge, MA, USA.
- Oliva M Section of Genetic Medicine, Department of Medicine, University of Chicago, Chicago, IL, USA.
- Muñoz-Aguirre M Centre for Genomic Regulation (CRG), The Barcelona Institute for Science and Technology, Barcelona, Catalonia, Spain.
- Kasela S New York Genome Center, New York, NY, USA.
- Wucher V Centre for Genomic Regulation (CRG), The Barcelona Institute for Science and Technology, Barcelona, Catalonia, Spain.
- Castel SE New York Genome Center, New York, NY, USA.
- Hamel AR The Broad Institute of MIT and Harvard, Cambridge, MA, USA.
- Viñuela A Department of Twin Research and Genetic Epidemiology, King's College London, London, UK.
- Roberts AL Department of Twin Research and Genetic Epidemiology, King's College London, London, UK.
- Mangul S Department of Computer Science, University of California, Los Angeles, Los Angeles, CA, USA.
- Wen X Department of Biostatistics, University of Michigan, Ann Arbor, MI, USA.
- Wang G Department of Human Genetics, University of Chicago, Chicago, IL, USA.
- Barbeira AN Section of Genetic Medicine, Department of Medicine, University of Chicago, Chicago, IL, USA.
- Garrido-Martín D Centre for Genomic Regulation (CRG), The Barcelona Institute for Science and Technology, Barcelona, Catalonia, Spain.
- Nadel BB Department of Molecular, Cellular, and Developmental Biology, University of California, Los Angeles, Los Angeles, CA, USA.
- Zou Y Department of Statistics, University of Chicago, Chicago, IL, USA.
- Bonazzola R Section of Genetic Medicine, Department of Medicine, University of Chicago, Chicago, IL, USA.
- Quan J Inflammation & Immunology, Pfizer, Cambridge, MA, USA.
- Brown A Department of Genetic Medicine and Development, University of Geneva Medical School, Geneva, Switzerland.
- Martinez-Perez A Unit of Genomic of Complex Diseases, Institut d'Investigació Biomèdica Sant Pau (IIB-Sant Pau), Barcelona, Spain.
- Soria JM Unit of Genomic of Complex Diseases, Institut d'Investigació Biomèdica Sant Pau (IIB-Sant Pau), Barcelona, Spain.
- Getz G Department of Genetic Medicine and Development, University of Geneva Medical School, Geneva, Switzerland.
- Dermitzakis ET Department of Twin Research and Genetic Epidemiology, King's College London, London, UK.
- Small KS Department of Human Genetics, University of Chicago, Chicago, IL, USA.
- Stephens M Foundational Neuroscience Center, AbbVie, Cambridge, MA, USA.
- Xi HS Section of Genetic Medicine, Department of Medicine, University of Chicago, Chicago, IL, USA.
- Im HK Centre for Genomic Regulation (CRG), The Barcelona Institute for Science and Technology, Barcelona, Catalonia, Spain.
- Guigó R The Broad Institute of MIT and Harvard, Cambridge, MA, USA.
- Segrè AV Section of Genetic Medicine, Department of Medicine, University of Chicago, Chicago, IL, USA.
- Stranger BE The Broad Institute of MIT and Harvard, Cambridge, MA, USA.
- Ardlie KG New York Genome Center, New York, NY, USA. skimhellmuth@gmail.com tlappalainen@nygenome.org.
- Lappalainen T
- 2020-09-11
Published in:
- Science (New York, N.Y.). - 2020
Cells
Gene Expression Regulation
Humans
Organ Specificity
Quantitative Trait Loci
RNA, Long Noncoding
Transcriptome
English
The Genotype-Tissue Expression (GTEx) project has identified expression and splicing quantitative trait loci in cis (QTLs) for the majority of genes across a wide range of human tissues. However, the functional characterization of these QTLs has been limited by the heterogeneous cellular composition of GTEx tissue samples. We mapped interactions between computational estimates of cell type abundance and genotype to identify cell type-interaction QTLs for seven cell types and show that cell type-interaction expression QTLs (eQTLs) provide finer resolution to tissue specificity than bulk tissue cis-eQTLs. Analyses of genetic associations with 87 complex traits show a contribution from cell type-interaction QTLs and enables the discovery of hundreds of previously unidentified colocalized loci that are masked in bulk tissue.
- Language
-
- English
- Open access status
- bronze
- Identifiers
-
- DOI 10.1126/science.aaz8528
- PMID 32913075
- Persistent URL
- https://folia.unifr.ch/global/documents/252138
Statistics
Document views: 67
File downloads:
- Full-text: 0